Introduction to Neural Networks
Data Science and Machine learning
Neural network jargon, that you will learn today…
- perceptron
- activation function
- epoch
- learning rate
- gradient descent
- backpropagation
- hidden layers
- deep learning
Introduction to Neural Networks
What is a Neural Network?
Neural networks are computational systems inspired by biological neural networks They can approximate non-linear relationships by learning from data.
They are composed of interconnected nodes (neurons) organized in layers.
Neural networks are particularly effective for:
Pattern recognition
Classification tasks
Regression problems
Applications: Remote Sensing and Image Recognition

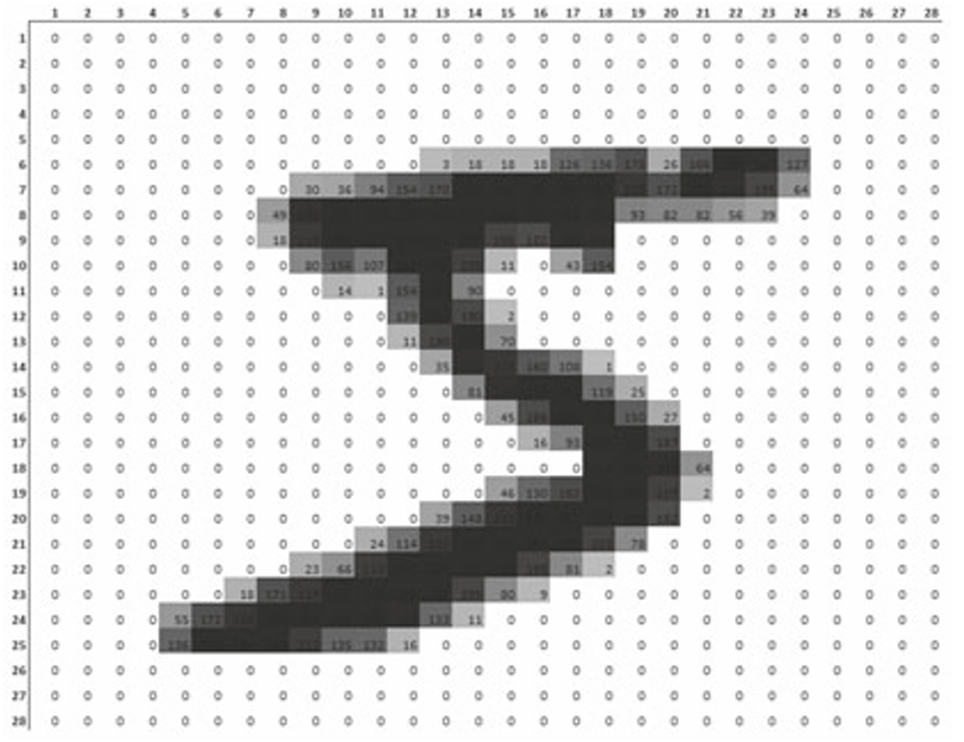
Applications: Image Recognition (I)

Applications: Image Recognition (II)

Applications: Macro (I)

Applications: Macro (II)

Applications (IV)
History (I)
McCulloch, W.S. and Pitts, W. (1943). A logical calculus of the ideas immanent in nervous activity. The bulletin of mathematical biophysics, 5(4):115–133.
History (II)
Rosenblatt, F. (1958). The perceptron: a probabilistic model for information storage and organization in the brain. Psychological review, 65(6), 386.
Main research questions:
- How do (highly developed) organisms store information?
- Does stored information influence the recognition of objects?
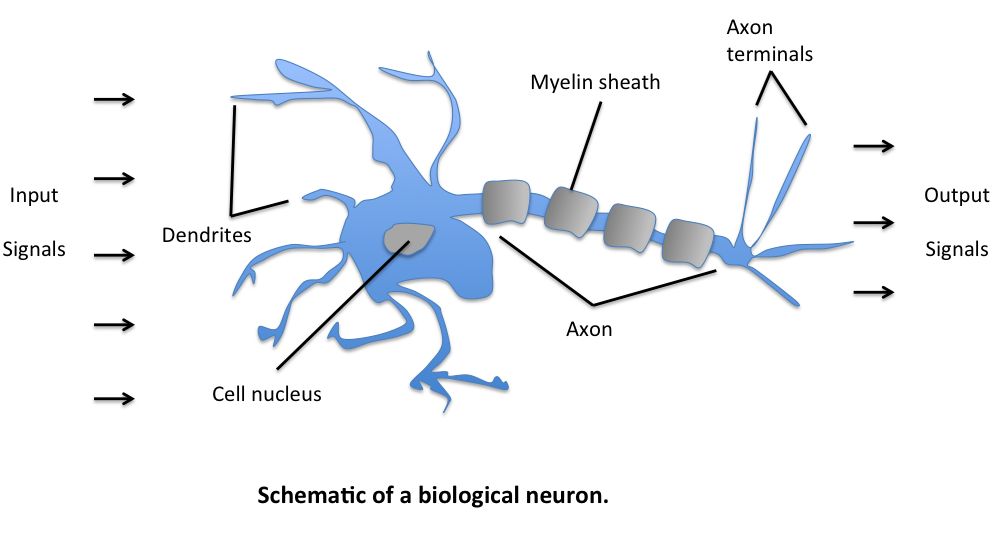
The biological neuron
Key insight: Learning occurs through strengthening or weakening synaptic connections
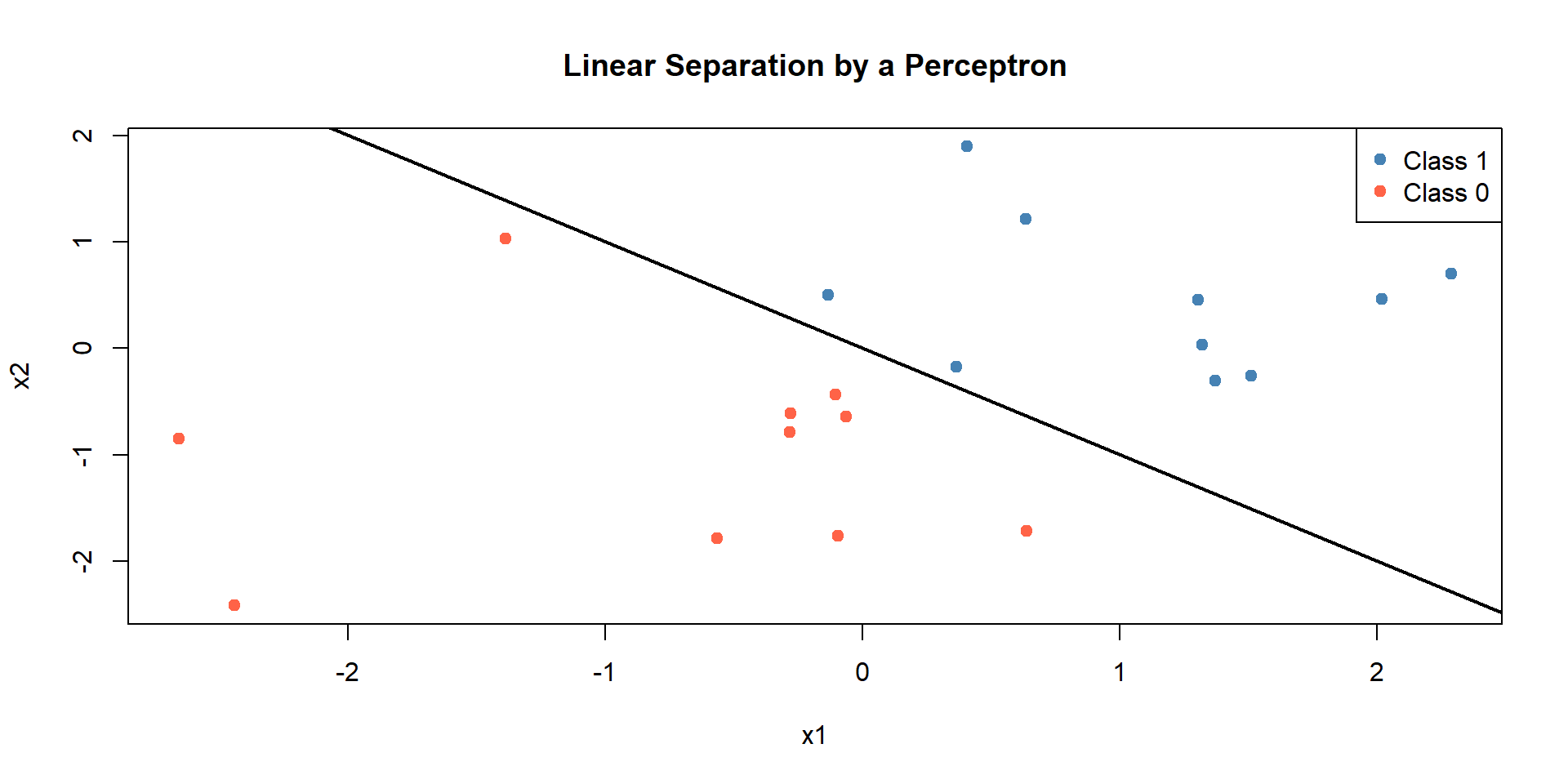
From neuron to perceptron
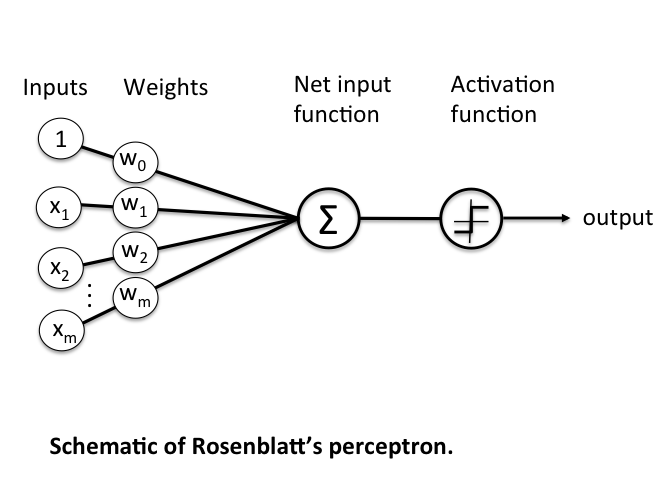
The perceptron is a mathematical model inspired by biological neurons.
Perceptron decision rule: main formula
\[ z = \sum_{i=1}^{n} w_i x_i + b \]
\[ \text{Output} = f(z) = \begin{cases} 1, & \text{if } z > 0 \\ 0, & \text{otherwise} \end{cases} \]
Example 1: Simple perceptron calculation
Given:
- \(w_1 = 0.5, w_2 = -0.4\)
- \(x_1 = 1, x_2 = 0\)
- \(b = -0.3\)
Calculation:
\[ z = (0.5 \times 1) + (-0.4 \times 0) + (-0.3) = 0.5 - 0.3 = 0.2 \]
Since \(z = 0.2 > 0\), output = 1
Example 2: Another perceptron calculation
Given:
- \(w_1 = 0.3, w_2 = 0.7\)
- \(x_1 = 1, x_2 = 1\)
- \(b = -0.5\)
Calculation:
\[ z = (0.3 \times 1) + (0.7 \times 1) + (-0.5) = 0.3 + 0.7 - 0.5 = 0.5 \]
Since \(z = 0.5 > 0\), output = 1
Example 3: Perceptron outputs 0
Given:
- \(w_1 = 0.2, w_2 = 0.3\)
- \(x_1 = 0\), \(x_2 = 1\)
- Bias \(b = -0.5\)
\[ z = (0.2 \times 0) + (0.3 \times 1) + (-0.5) = 0.3 - 0.5 = -0.2 \]
Since \(z = -0.2 < 0\), output = 0
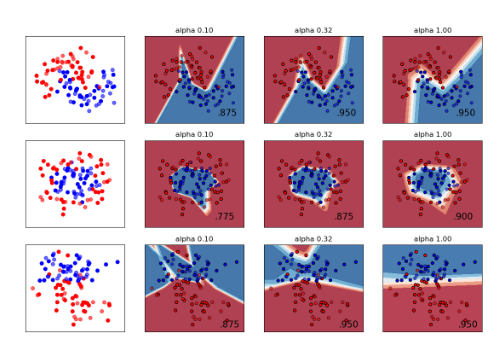
Example 1: Simple XOR Problem
XOR Problem
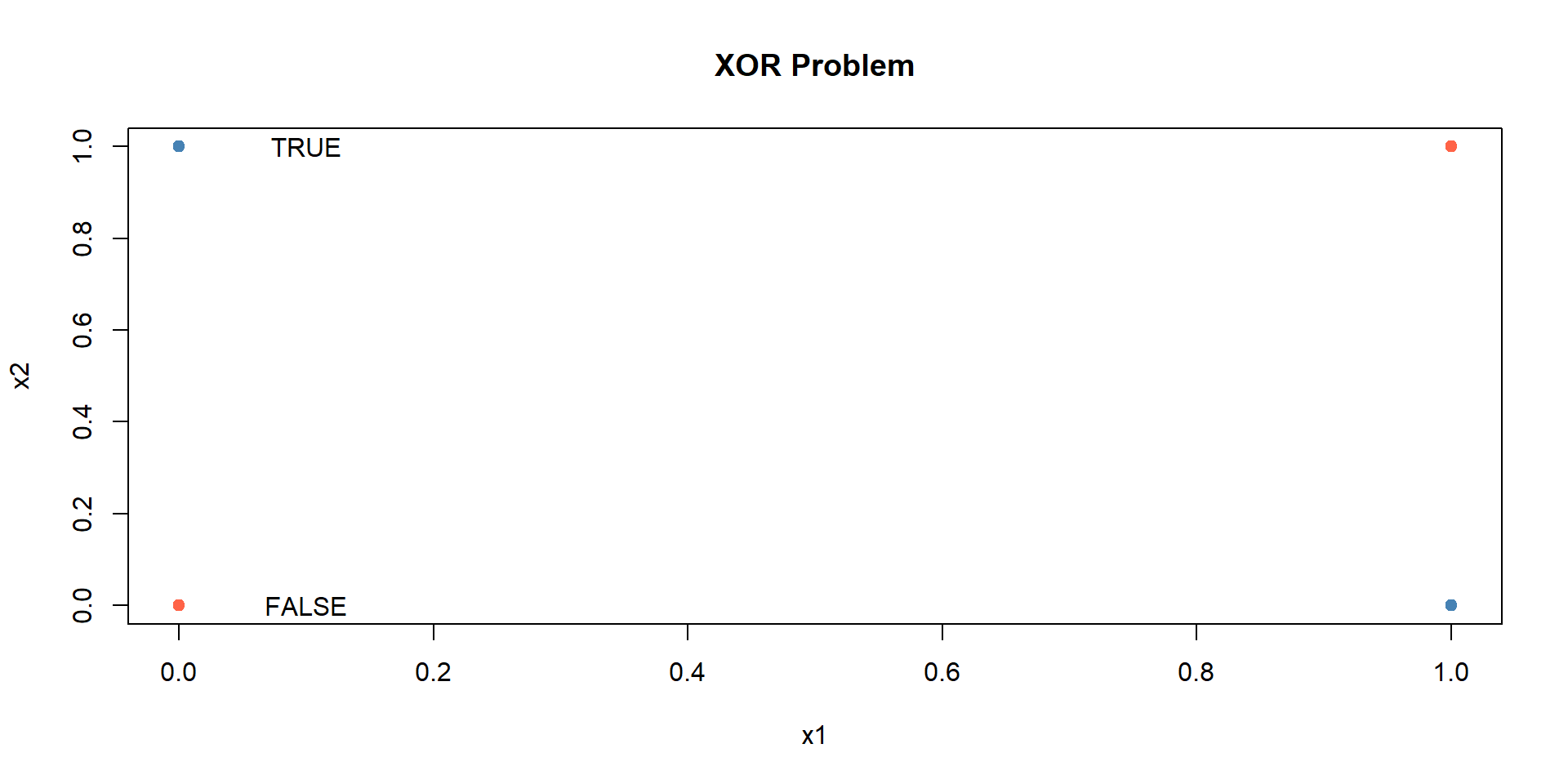
Non-linear classification with neural networks
Neural Network Architecture
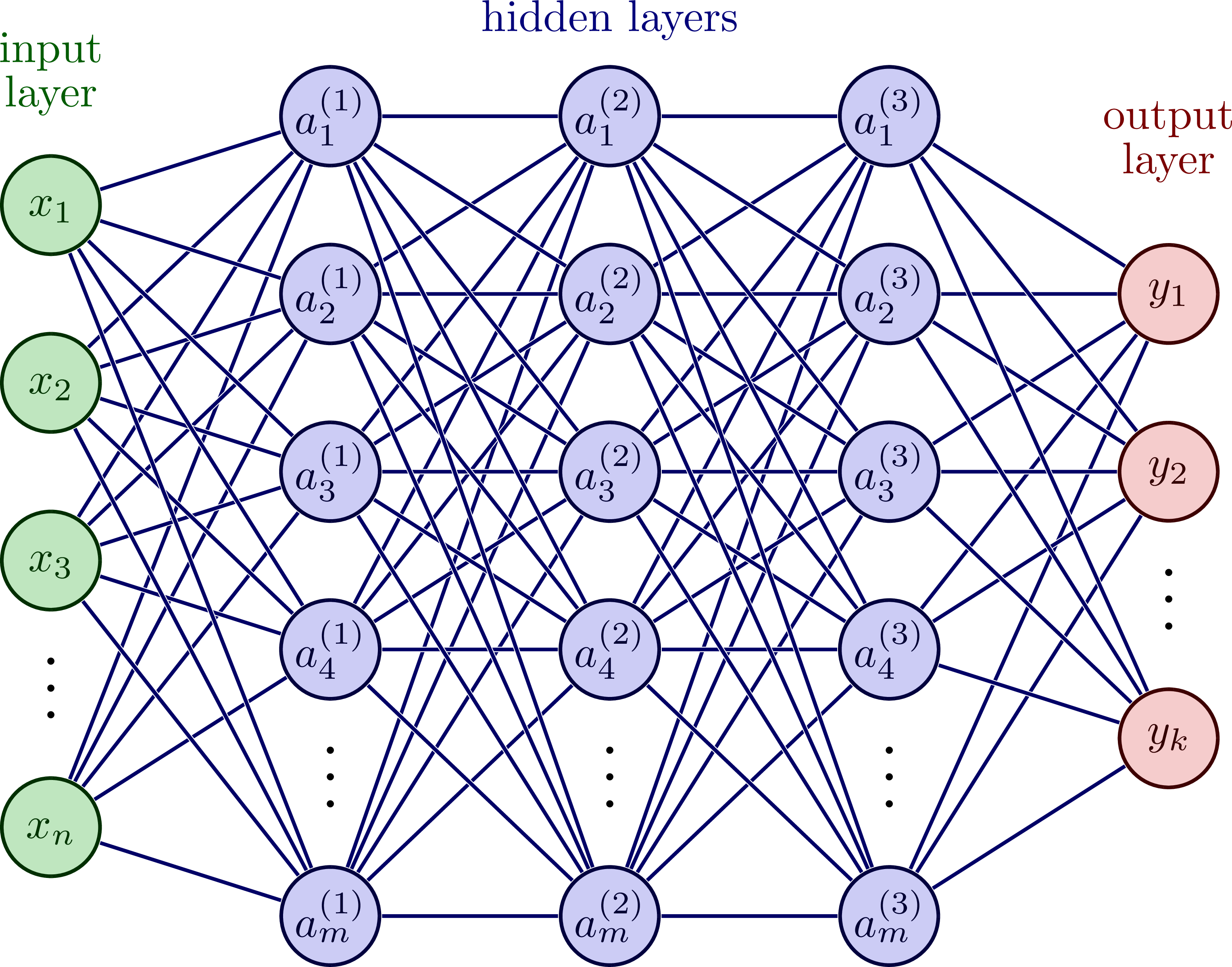
Basic Components of a Neural Network (I)
A single neuron performs:
\[z = \sum_{i=1}^{n} w_i x_i + b\]
Where:
- \(x_i\) = input values
- \(w_i\) = weights (connection strengths)
- \(b\) = bias term
Basic Components of a Neural Network (II)
\[y = f(z)\]
\(f\) = activation function
\(y\) = output
Activation Functions
Activation Functions
Activation functions introduce non-linearity to the network.
- Without activation functions, neural networks would be linear models
- Activation functions enable networks to learn complex patterns
Linear function
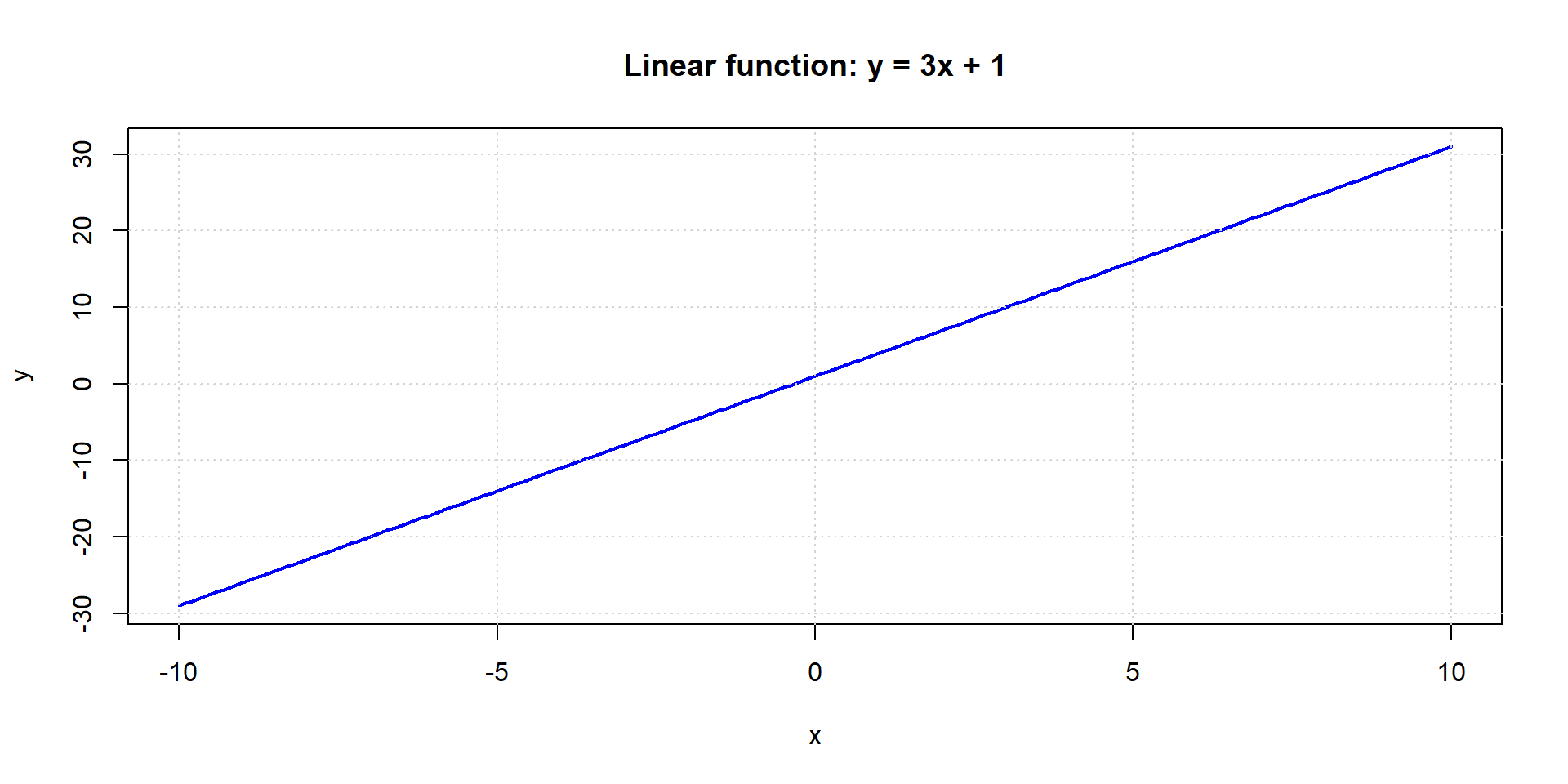
Higher degree / non-linear functions
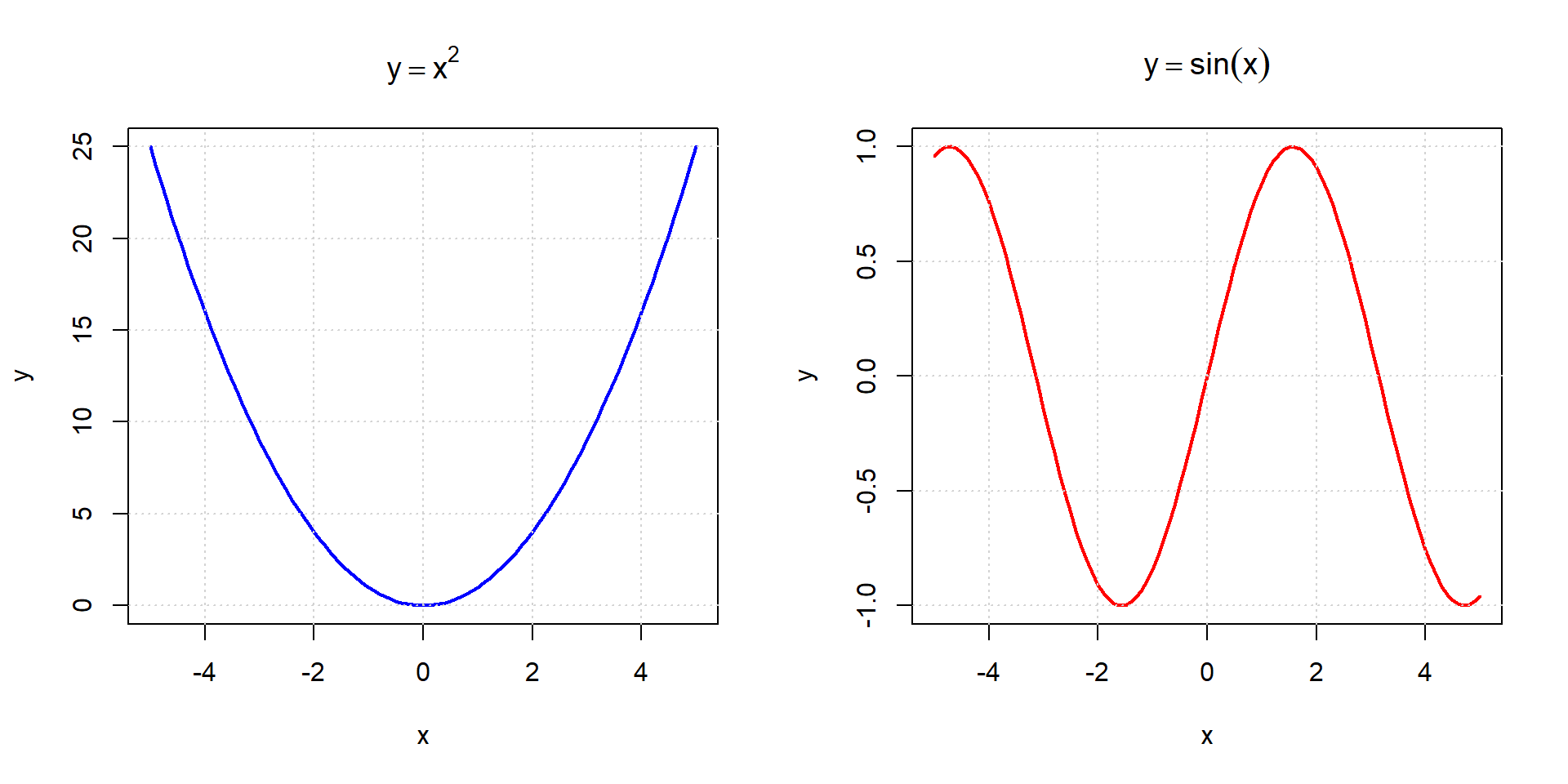
Step function
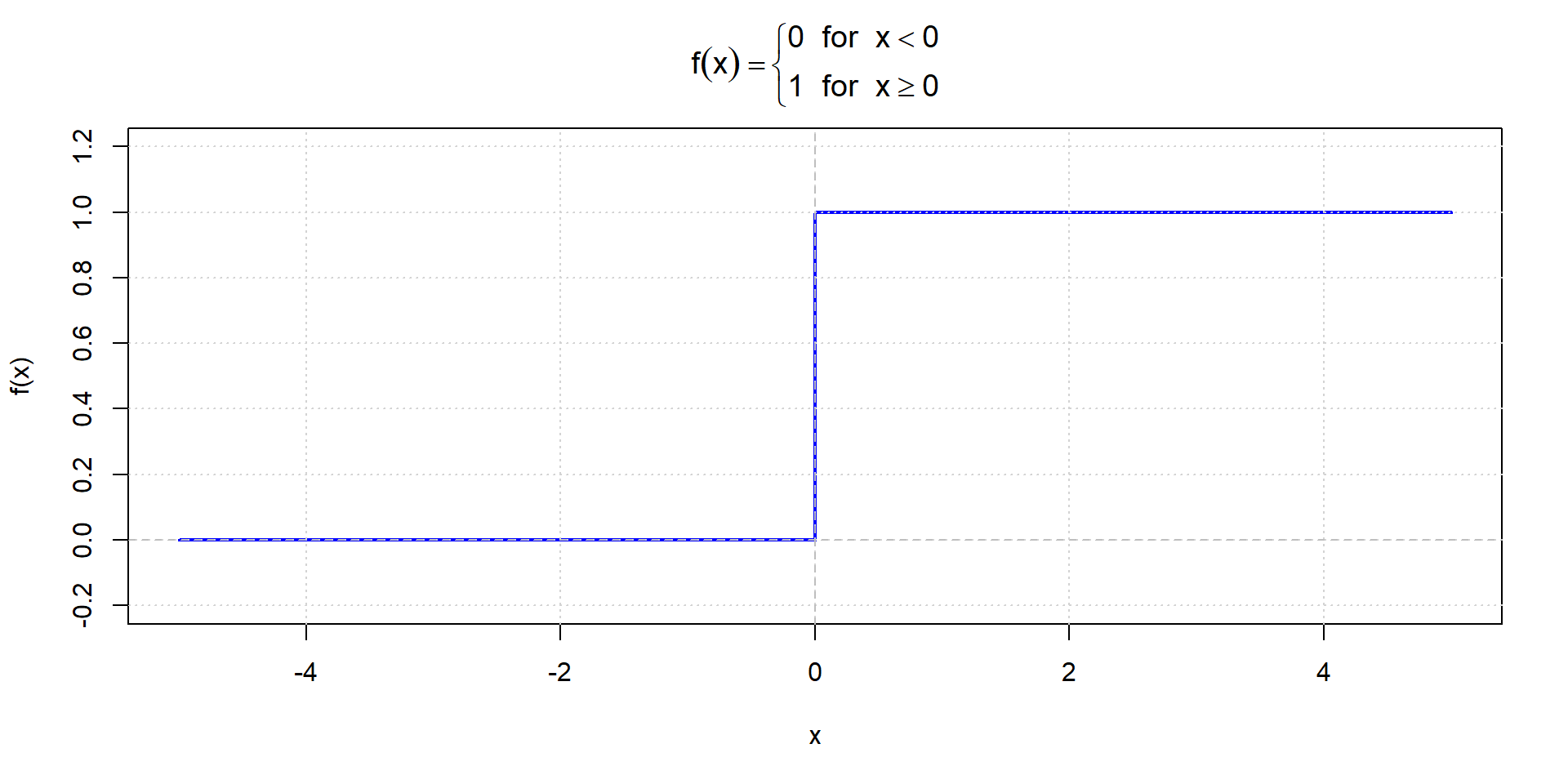
Sigmoid function
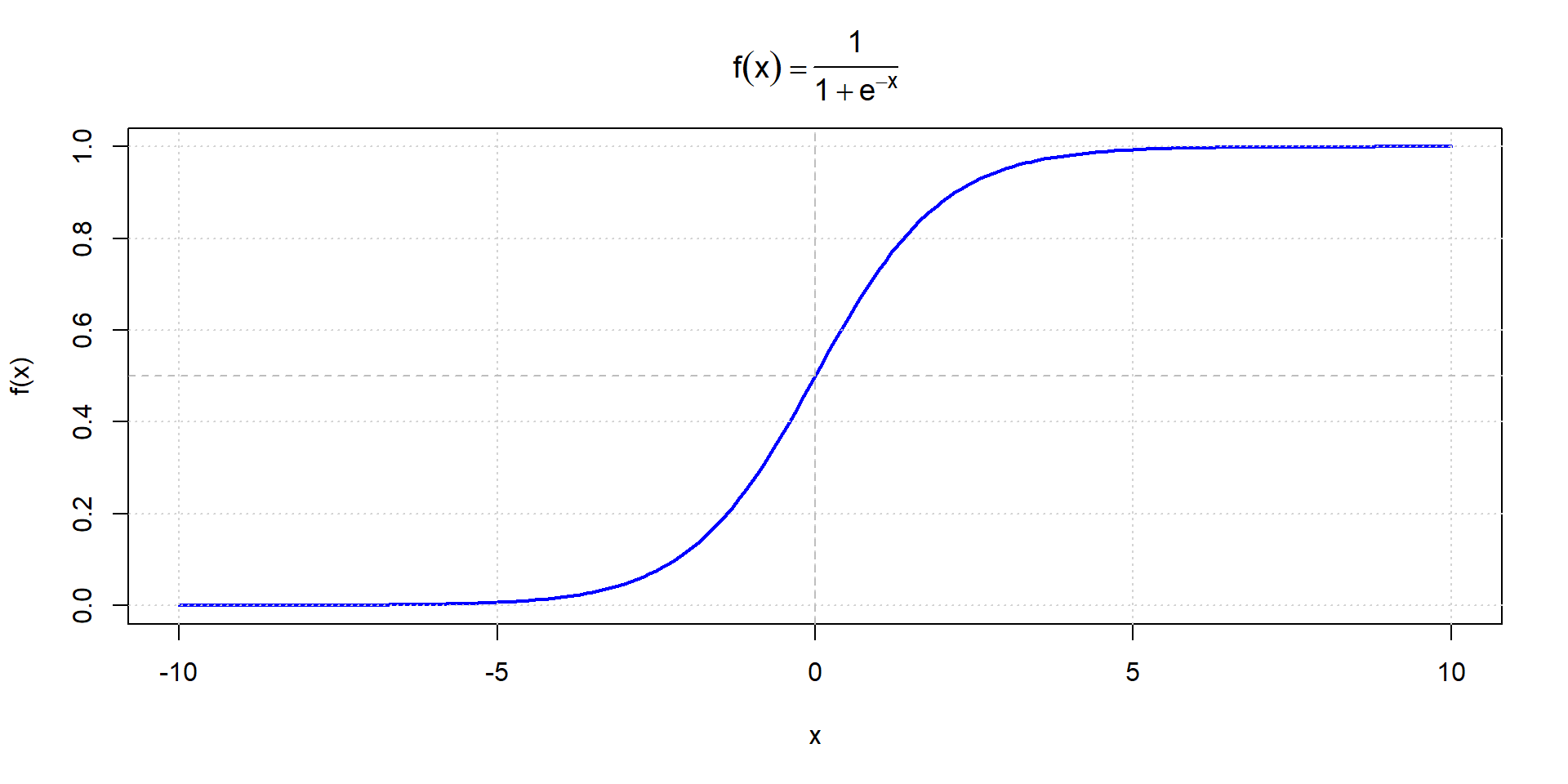
Hyperbolic tangent (tanh)
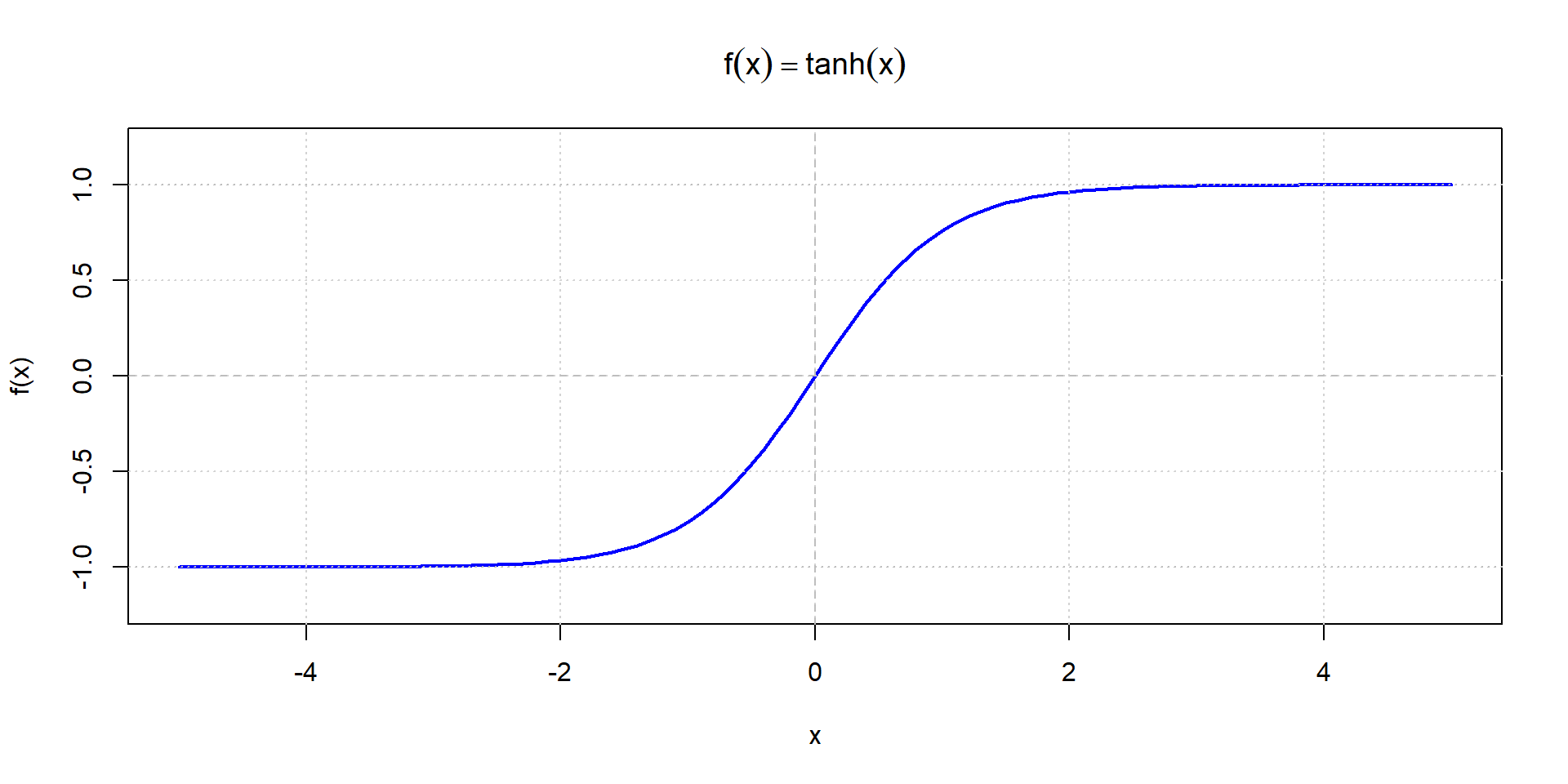
Sigmoid vs. tanh comparison
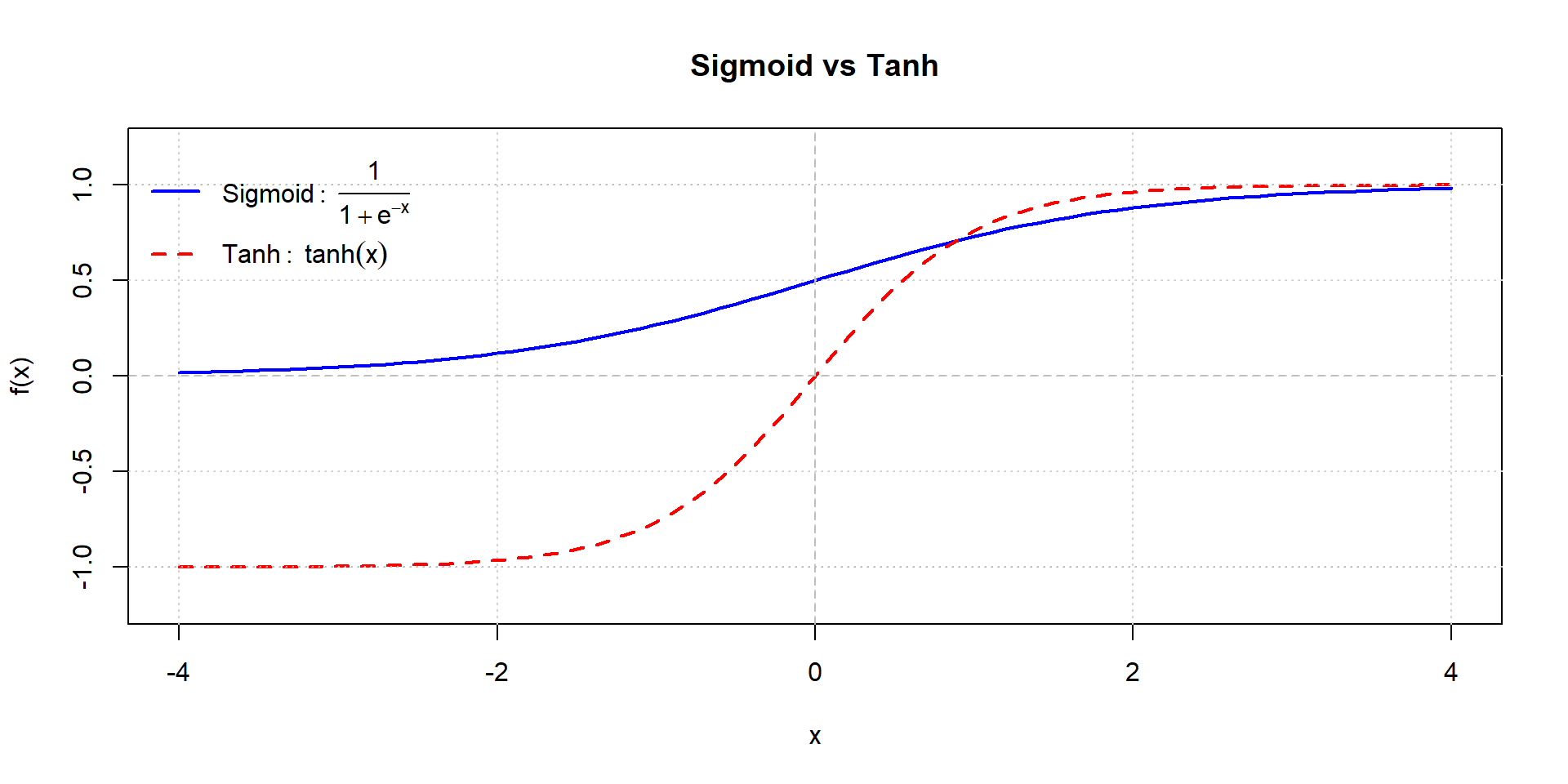
ReLU function
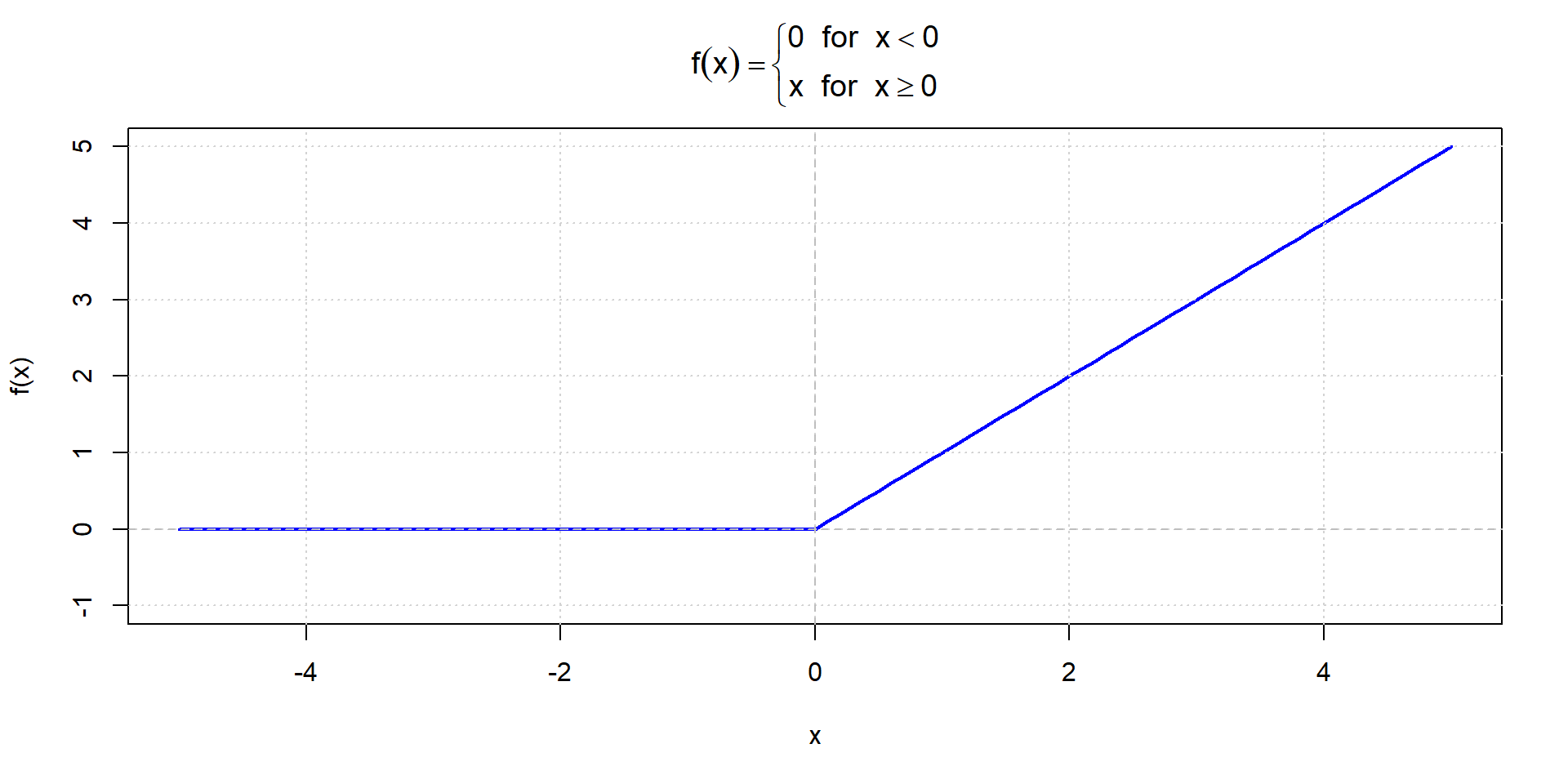
Training Neural Networks
What is Training
“Training is the act of presenting the network with some sample data and modifying the weights to better approximate the desired function”
- The network learns by adjusting weights and biases
- Goal: Minimize the difference between predicted and actual outputs
Forward Propagation and Backpropagation
Forward Propagation
Definition: Processing from input layer → hidden layer → output layer
The forward pass computes:
- Weighted sum at each neuron: \(z = wx + b\)
- Apply activation function: \(a = f(z)\)
- Pass to next layer
- Repeat until output layer
- Calculate prediction
Forward Propagation Steps
For a network with one hidden layer:
Hidden Layer: \[z^{(1)} = w^{(1)}x + b^{(1)}\] \[a^{(1)} = f(z^{(1)})\]
Output Layer: \[z^{(2)} = w^{(2)}a^{(2)} + b^{(2)}\] \[\hat{y} = f(z^{(2)})\]
Computing the Error
- At the output layer, we compute the error:
\[\text{Error} = \text{Predicted Output} - \text{Actual Output}\]
Backpropagation (I)
Definition: The process of propagating the error backward through the network to update weights
- We correct the error in the process of backpropagation
- Uses the partial derivative of each neuron’s activation function
- The amount by which the weight should change is determined by gradient descent
Backpropagation (II)
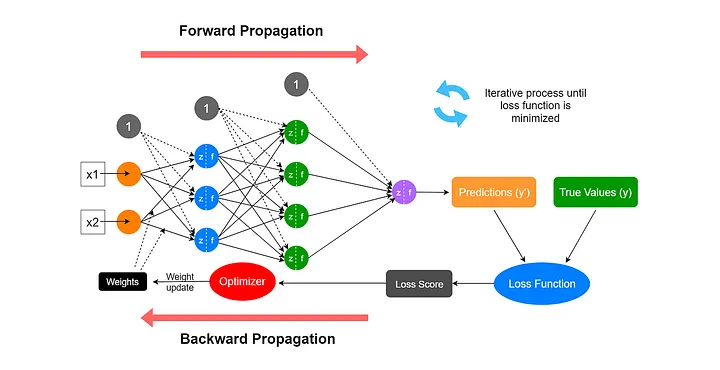
The Training Algorithm (I)
Complete training process:
- Take the matrix-based input
- Assign initially random weights and biases
- Apply inputs to the network
- Calculate the output through the hidden layer(s) (forward propagation)
The Training Algorithm (II)
- At the output layer: calculate the error
- Compute error signals for previous layers using the partial derivative of the activation function (backpropagation)
- Use the error signals to compute weight adjustments
- Apply the weight adjustments
- Repeat steps 3 to 8 until error is minimized
Gradient Descent
What is Gradient Descent?
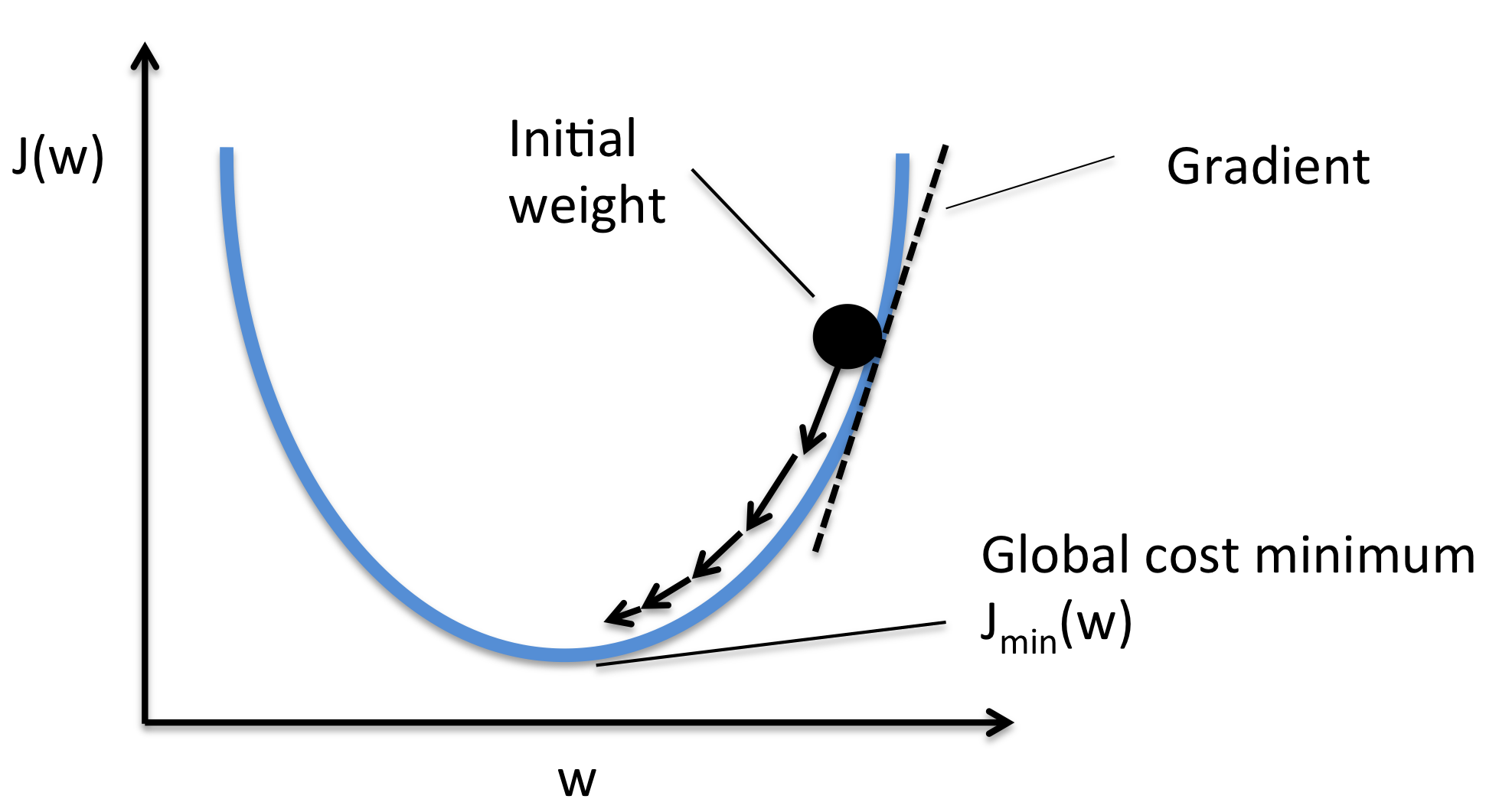
- Optimization algorithm to minimize the loss function
- Iteratively adjusts weights in the direction that reduces error
What is Gradient Descent?
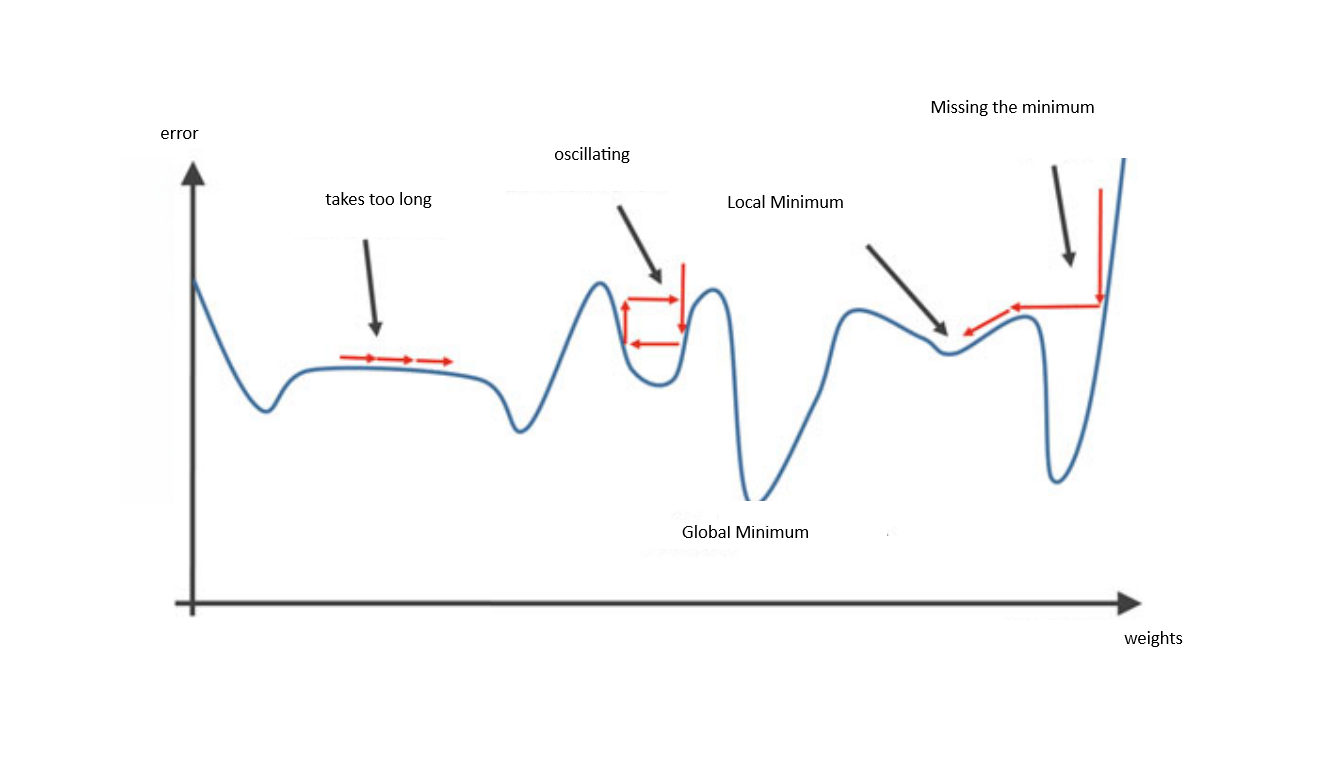
The Learning Rate
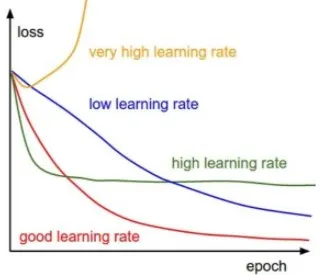
The learning rate is our step-size in adjusting weights:
New weight = Existing weight — Learning rate * Gradient
Gradient Descent explained
3Blue1Brown Video
“Gradient descent, how neural networks learn” Watch on YouTube
Multi-layer Neural Network Structure

Fully connected: each neuron connects to all neurons in next layer
Multiple Hidden Layers (Deep Learning)
Shallow networks: 1 hidden layer
Deep networks: 2+ hidden layers
Deeper networks can learn more complex patterns
Trade-offs:
- More expressive power
- Longer training time
- Requires more data
Types of Learning
Three main paradigms:
- Supervised Learning
- Unsupervised Learning
- Reinforcement Learning
Supervised Learning
- Training with labeled data (input-output pairs)
- The network learns the mapping from inputs to outputs
- Examples:
- Image classification (cat vs. dog)
- Spam detection
- House price prediction
- Most common type for neural networks
Unsupervised Learning
- Training with unlabeled data
- The network discovers patterns and structures
- Examples:
- Customer segmentation
- Anomaly detection
- No predefined target values
Reinforcement Learning
- Learning through trial and error
- Agent receives rewards or penalties for actions
- Goal: Maximize cumulative reward over time
- Examples:
- Game playing
- Robotics
Some Training Concepts
Epoch
Definition: One complete pass through the entire training dataset
- One iteration of providing input and updating network weights
- Training typically requires multiple epochs
- Each epoch:
- All training examples are seen once
- Weights are updated multiple times
- Model progressively improves
Epoch vs Batch vs Iteration
- Batch: Subset of training data processed together
- Iteration: One weight update (processing one batch)
- Epoch: Complete pass through all data
Example with 1000 samples:
Batch size = 100 → 10 iterations per epoch
Batch size = 250 → 4 iterations per epoch
Choosing Number of Epochs
- Too few epochs: Underfitting
- Too many epochs: Overfitting
Scaling
Why Scale Your Data?
Neural networks require scaled input data:
- Features often have different ranges (e.g., 0-1 vs 1000-10000)
- Large values can dominate gradients
- Leads to:
- Slow convergence
- Unstable training
- Poor performance
Scaling Methods
1. Min-Max Normalization (scales to 0-1):
\[x_{\text{norm}} = \frac{x - x_{\min}}{x_{\max} - x_{\min}}\]
2. Z-score Standardization (mean=0, std=1):
\[x_{\text{std}} = \frac{x - \mu}{\sigma}\]
Understanding the Neural Network
Example: Boston Dataset
The Boston Dataset (I)
The Boston data frame contains 506 rows and 14 columns of housing market data from the Boston area.
Prediction goal: medv is our response variable (= median value of owner-occupied homes in USD thousand). We fit a regression model to explain variation in owner-occupied home value.
Variable Descriptions
Left Column
crim: Per capita crime rate by town
zn: Proportion of residential land zoned for lots over 25,000 sq.ft.
indus: Proportion of non-retail business acres per town
chas: Charles River dummy variable (= 1 if tract bounds river; 0 otherwise)
nox: Nitrogen oxides concentration (parts per 10 million)
rad: Index of accessibility to radial highways
tax: Full-value property-tax rate per $10,000
Right Column
rm: Average number of rooms per dwelling
age: Proportion of owner-occupied units built prior to 1940
dis: Weighted mean of distances to five Boston employment centres
ptratio: Pupil-teacher ratio by town
black: \(1000(B_k - 0.63)^2\) where \(B_k\) is the proportion of blacks by town
lstat: Lower status of the population (percent)
medv: Median value of owner-occupied homes in $1,000s (TARGET)
The Boston Dataset (II)
#|echo: true
library("neuralnet")
library("MASS")
library("dplyr")
library("mlbench")
library(magrittr)
library(patchwork)
library(ggplot2)
library("reshape2")
set.seed("3456")
data = Boston
head(data) crim zn indus chas nox rm age dis rad tax ptratio black lstat
1 0.00632 18 2.31 0 0.538 6.575 65.2 4.0900 1 296 15.3 396.90 4.98
2 0.02731 0 7.07 0 0.469 6.421 78.9 4.9671 2 242 17.8 396.90 9.14
3 0.02729 0 7.07 0 0.469 7.185 61.1 4.9671 2 242 17.8 392.83 4.03
4 0.03237 0 2.18 0 0.458 6.998 45.8 6.0622 3 222 18.7 394.63 2.94
5 0.06905 0 2.18 0 0.458 7.147 54.2 6.0622 3 222 18.7 396.90 5.33
6 0.02985 0 2.18 0 0.458 6.430 58.7 6.0622 3 222 18.7 394.12 5.21
medv
1 24.0
2 21.6
3 34.7
4 33.4
5 36.2
6 28.7Scaling
We first scale the data:
\[x_{\text{scaled}} = \frac{x - x_{\min}}{x_{\max} - x_{\min}}\]
crim zn indus chas
Min. : 0.00632 Min. : 0.00 Min. : 0.46 Min. :0.00000
1st Qu.: 0.08205 1st Qu.: 0.00 1st Qu.: 5.19 1st Qu.:0.00000
Median : 0.25651 Median : 0.00 Median : 9.69 Median :0.00000
Mean : 3.61352 Mean : 11.36 Mean :11.14 Mean :0.06917
3rd Qu.: 3.67708 3rd Qu.: 12.50 3rd Qu.:18.10 3rd Qu.:0.00000
Max. :88.97620 Max. :100.00 Max. :27.74 Max. :1.00000
nox rm age dis
Min. :0.3850 Min. :3.561 Min. : 2.90 Min. : 1.130
1st Qu.:0.4490 1st Qu.:5.886 1st Qu.: 45.02 1st Qu.: 2.100
Median :0.5380 Median :6.208 Median : 77.50 Median : 3.207
Mean :0.5547 Mean :6.285 Mean : 68.57 Mean : 3.795
3rd Qu.:0.6240 3rd Qu.:6.623 3rd Qu.: 94.08 3rd Qu.: 5.188
Max. :0.8710 Max. :8.780 Max. :100.00 Max. :12.127
rad tax ptratio black
Min. : 1.000 Min. :187.0 Min. :12.60 Min. : 0.32
1st Qu.: 4.000 1st Qu.:279.0 1st Qu.:17.40 1st Qu.:375.38
Median : 5.000 Median :330.0 Median :19.05 Median :391.44
Mean : 9.549 Mean :408.2 Mean :18.46 Mean :356.67
3rd Qu.:24.000 3rd Qu.:666.0 3rd Qu.:20.20 3rd Qu.:396.23
Max. :24.000 Max. :711.0 Max. :22.00 Max. :396.90
lstat medv
Min. : 1.73 Min. : 5.00
1st Qu.: 6.95 1st Qu.:17.02
Median :11.36 Median :21.20
Mean :12.65 Mean :22.53
3rd Qu.:16.95 3rd Qu.:25.00
Max. :37.97 Max. :50.00 Build the training dataset
split <- sample(1:nrow(data),round(0.75*nrow(data)))
train_data <- as.data.frame(data_scaled[split,])
test_data <- as.data.frame(data_scaled[-split, ])
nrow(test_data)[1] 126[1] 380 crim zn indus chas
Min. :0.0000522 Min. :0.0000 Min. :0.01026 Min. :0.00000
1st Qu.:0.0008616 1st Qu.:0.0000 1st Qu.:0.17815 1st Qu.:0.00000
Median :0.0033157 Median :0.0000 Median :0.34604 Median :0.00000
Mean :0.0398760 Mean :0.1092 Mean :0.39591 Mean :0.07105
3rd Qu.:0.0463340 3rd Qu.:0.1250 3rd Qu.:0.64663 3rd Qu.:0.00000
Max. :0.8264345 Max. :1.0000 Max. :1.00000 Max. :1.00000
nox rm age dis
Min. :0.0000 Min. :0.05787 Min. :0.0000 Min. :0.0000
1st Qu.:0.1296 1st Qu.:0.44309 1st Qu.:0.4343 1st Qu.:0.0890
Median :0.3148 Median :0.50326 Median :0.7626 Median :0.1952
Mean :0.3516 Mean :0.51985 Mean :0.6737 Mean :0.2432
3rd Qu.:0.5062 3rd Qu.:0.58392 3rd Qu.:0.9323 3rd Qu.:0.3648
Max. :1.0000 Max. :1.00000 Max. :1.0000 Max. :0.8712
rad tax ptratio black
Min. :0.0000 Min. :0.001908 Min. :0.0000 Min. :0.005547
1st Qu.:0.1304 1st Qu.:0.179389 1st Qu.:0.5106 1st Qu.:0.940932
Median :0.1739 Median :0.272901 Median :0.6809 Median :0.986321
Mean :0.3926 Mean :0.434281 Mean :0.6259 Mean :0.890165
3rd Qu.:1.0000 3rd Qu.:0.914122 3rd Qu.:0.8085 3rd Qu.:0.998260
Max. :1.0000 Max. :1.000000 Max. :1.0000 Max. :1.000000
lstat medv
Min. :0.02042 Min. :0.0000
1st Qu.:0.15542 1st Qu.:0.2567
Median :0.27525 Median :0.3578
Mean :0.30949 Mean :0.3872
3rd Qu.:0.42557 3rd Qu.:0.4383
Max. :1.00000 Max. :1.0000 Set-up the neural net
Neural Net structure
Take predictions, report the error
# predict with neural net on test data
predict_net <- predict(net, test_data[,1:13])
predict_net_unscaled <- as.data.frame(predict_net * (max(data$medv)- min(data$medv)) + min (data$medv))
# compute RMSEs
rmse.net <- caret::RMSE(predict_net, test_data$medv)
print(paste("RMSE neural net:", rmse.net))[1] "RMSE neural net: 0.0631425251809264"test_unscaled <- as.data.frame((test_data$medv) * (max(data$medv)- min(data$medv)) + min (data$medv))
# Compare distribution of actual and predicted testing dataset
summary(predict_net_unscaled) V1
Min. : 7.443
1st Qu.:17.554
Median :21.831
Mean :22.881
3rd Qu.:26.953
Max. :54.213 (test_data$medv) * (max(data$medv) - min(data$medv)) + min(data$medv)
Min. : 5.00
1st Qu.:17.80
Median :21.80
Mean :22.85
3rd Qu.:27.38
Max. :50.00 Regression model
Call:
glm(formula = medv ~ ., data = train_data)
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.506041 0.064010 7.906 3.17e-14 ***
crim -0.201157 0.090682 -2.218 0.02715 *
zn 0.103714 0.037553 2.762 0.00604 **
indus -0.012132 0.046426 -0.261 0.79399
chas 0.070491 0.022767 3.096 0.00211 **
nox -0.226284 0.049973 -4.528 8.06e-06 ***
rm 0.435719 0.058404 7.460 6.33e-13 ***
age -0.005258 0.033573 -0.157 0.87564
dis -0.405556 0.059894 -6.771 5.10e-11 ***
rad 0.135291 0.042400 3.191 0.00154 **
tax -0.107243 0.055410 -1.935 0.05370 .
ptratio -0.193583 0.033832 -5.722 2.19e-08 ***
black 0.087407 0.027217 3.211 0.00144 **
lstat -0.420892 0.049694 -8.470 6.06e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for gaussian family taken to be 0.01192427)
Null deviance: 16.5010 on 379 degrees of freedom
Residual deviance: 4.3643 on 366 degrees of freedom
AIC: -588.96
Number of Fisher Scoring iterations: 2Compare RMSE
predict_lm <- predict(Regression_Model, test_data[,1:13])
predict_lm_unscaled <- as.data.frame(predict_lm * (max(data$medv)- min(data$medv)) + min (data$medv))
rmse.lm <- caret::RMSE(predict_lm, test_data$medv)
# Print the RMSE
print(paste("Root Mean Squared Error Linear Model:", rmse.lm))[1] "Root Mean Squared Error Linear Model: 0.0951616561991058"[1] "Root Mean Squared Error Neural Net: 0.0631425251809264"Plot actual predictions on tested data (I)
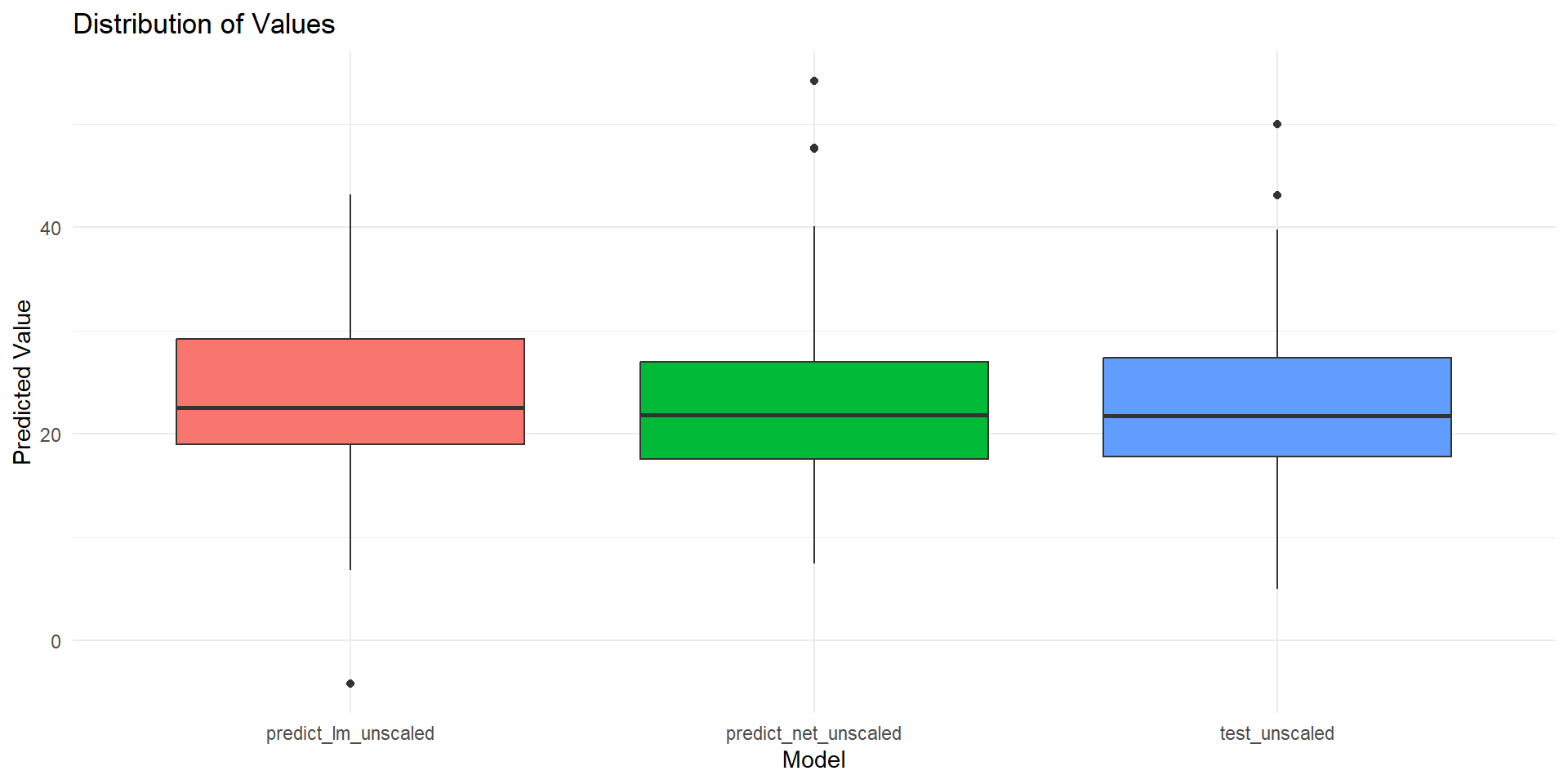
Plot actual predictions on tested data (II)
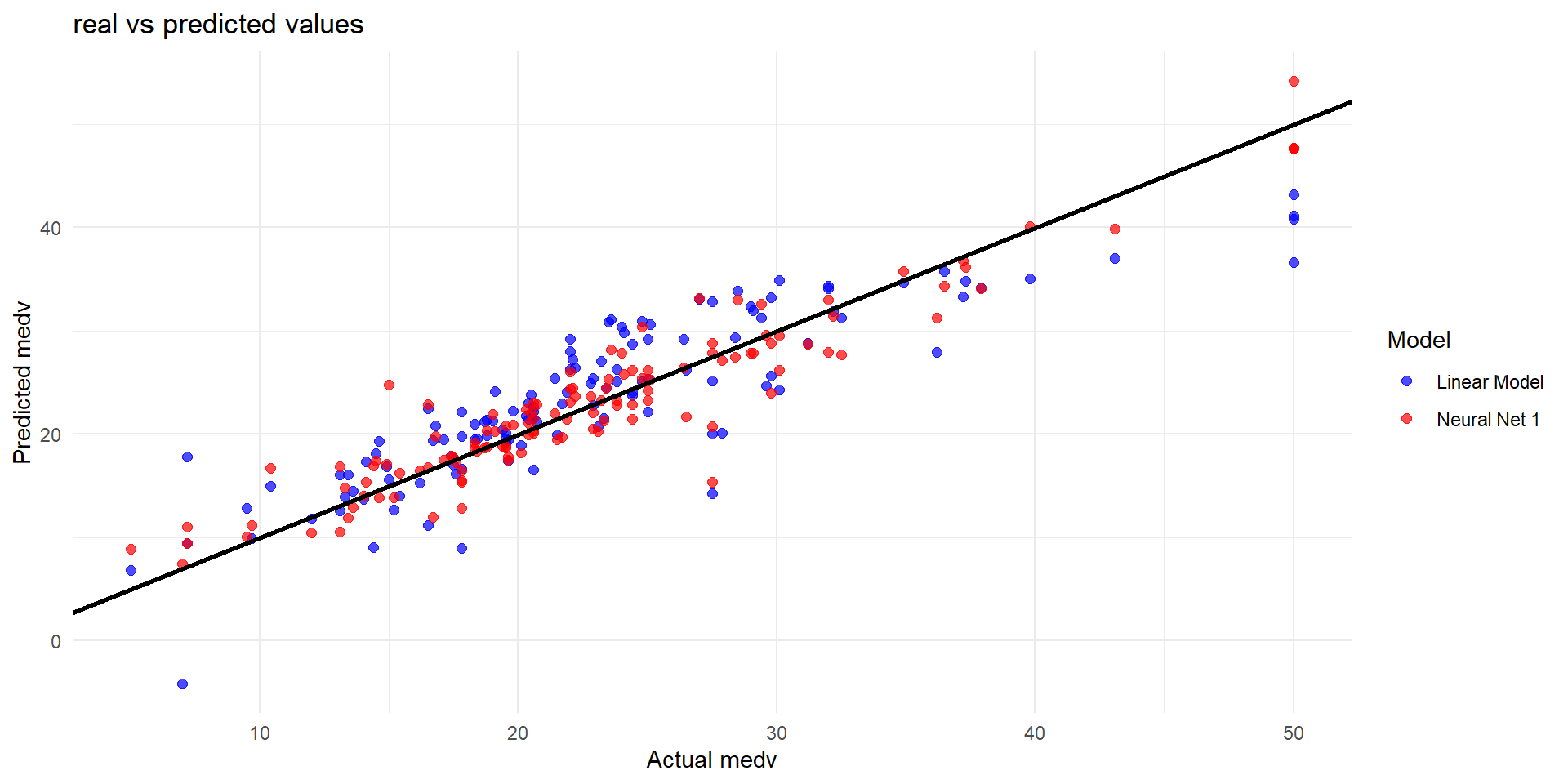
Key Takeaways
- Neural networks learn by adjusting weights and biases
- Forward propagation: Compute predictions
- Backpropagation: Compute gradients and update weights
- Gradient descent: Optimization algorithm
- Epochs: Multiple passes through data needed
- Data scaling: Essential preprocessing step
- Three learning types: Supervised, Unsupervised, Reinforcement
Disadvantages of Neural Networks
NN are black-boxes
NN and other machine learning tools are estimators - they cannot replace causal identification
NN are computationally expensive and time-intensive
there is no specific rulebook on parameter settings
global vs local optimum
Assigment: Estimation exercise
Take a dataset of your choice and a dependent variable of interest
Explore the dataset: Set-up some descriptives (plot or table) for your variables
Set-up a prediction exercise:
Choose a Machine Learning Tool of your choice and predict your dependent variable
Based on the same data, use OLS
Compare the results from OLS and your Machine Learning base estimator using RMSE or other performance criteria
What of these performed better? What do you find more suitable for your data source?
References and further readings
- Ciaburro, G., & Venkateswaran, B. (2017). Neural Networks with R. Packt Publishing.
- Goodfellow, I., Bengio, Y., & Courville, A. (2016). Deep Learning. MIT Press.
- 3Blue1Brown. (2017). Neural Networks Series. YouTube.
- Sonnet, D., Perceptron, V., & Learning, D. (2022). Neuronale Netze kompakt. IT kompakt. https://doi. org/10.1007/978-3-658-29081-8.
- Zhang Z. Neural networks: further insights into error function, generalized weights and others. Ann Transl Med. 2016 Aug;4(16):300. doi: 10.21037/atm.2016.05.37. PMID: 27668220; PMCID: PMC5009026.